Get Started¶
Concepts¶
The SCIPR algorithm for aligning scRNA-seq data is an iterative algorithm with the goal of learning a global function which can be used to align cells of one type of batch to another type of batch. It proceeds in steps, each of which consists of two phases:
- Matching - pairing cells between two batches of data (i.e. cells which have the same “state”).
- Transforming - learning a function to move the cells of one batch closer to their assigned counterpart (the cell they are paired with from step (1)) in the other batch.
Phase 1 corresponds to panel 2) in the figure below, and phase 2 corresponds to panel 3):
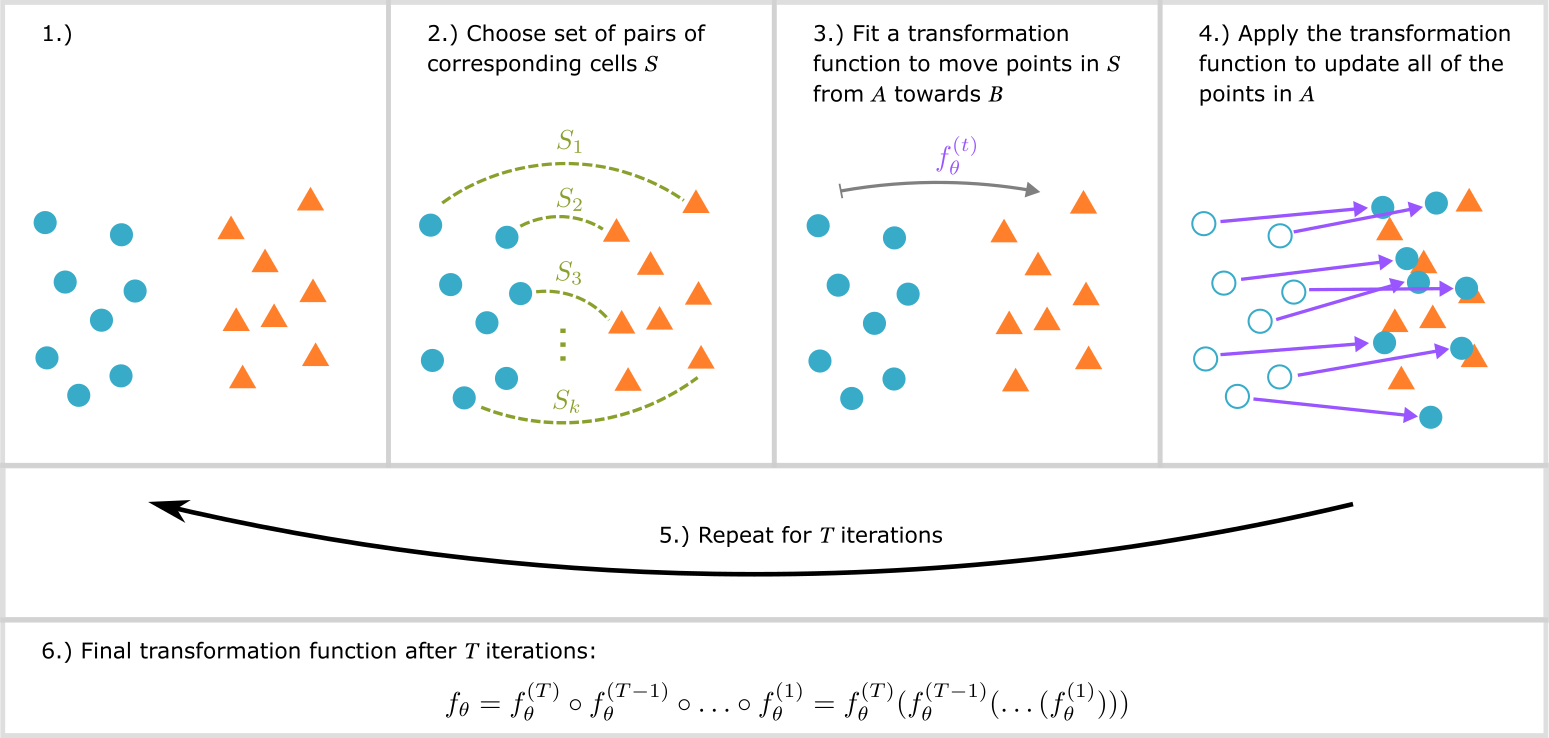
Finally, after all of the steps of SCIPR, the resulting transformation function is actually the composition of the learned functions at each step (panel 6) in the figure above).
Implementation¶
The SCIPR algorithm to learn the transformation function is implemented as the
scipr.SCIPR.fit() method. Users can secify which algorithms to use for
the matching and the transforming, either from among the options we provide in
our API, or by implementing their own custom extensions.
In particular, scipr implements the Strategy design pattern [GHJV94], where
each matching or transformation algorithm is implemented as a concrete subclass
of either the abstract scipr.matching.Match or
scipr.transform.Transformer classes.
Examples¶
Use the classic Iterative Closest Points (ICP) [BeMc92] algorithm to align cells:
import scipr
from scipr.matching import Closest
from scipr.transform import Rigid
closest = Closest()
rigid = Rigid()
model = scipr.SCIPR(match_algo=closest,
transform_algo=rigid,
input_normalization='l2')
model.fit(batch_A, batch_B)
# Apply the model to get the aligned result
batch_A_aligned = model.trasform(batch_A)
Alternatively, we recommend trying matching and transformation functions more suited to scRNA-seq data:
from scipr.matching import MNN
from scipr.transform import Affine
mnn = MNN()
affine = Affine()
model = scipr.SCIPR(match_algo=mnn,
transform_algo=affine,
input_normalization='l2')
model.fit(batch_A, batch_B)
batch_A_aligned = model.trasform(batch_A)
Logging¶
scipr uses Python’s built-in logging module to report informational messages and training metrics (e.g. loss values at each gradient descent iteration), and so normally you should see nothing printed to stdout (unless it’s warning message). If you would like to see this informational text, configure the Python logger with the appropriate level like so:
import logging
logging.basicConfig(level=logging.INFO)
TensorBoard¶
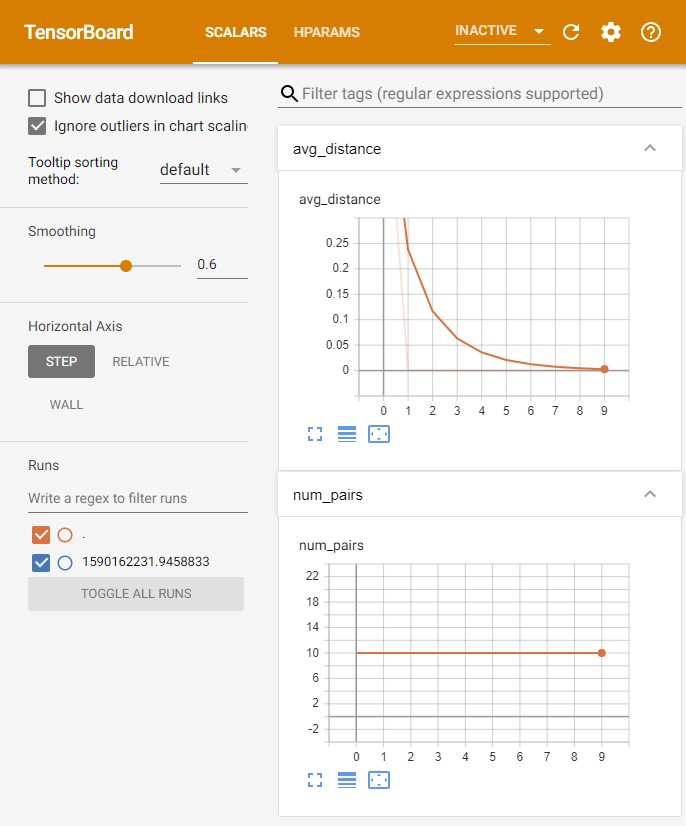
Some scipr functions can be enabled with TensorBoard logging, giving you a useful “dashboard” view of model metrics as the SCIPR model is fit, as in the example screenshot below. It can be very useful for debugging an alignment. See Installation for enabling this feature.
References
| [GHJV94] | Gamma, E., Helm, R., Johnson, R., & Vlissides, J. (1994). Design Patterns: Elements of Reusable Object-Oriented Software. |
| [BeMc92] | Besl, P. J., & McKay, H. D. (1992). A method for registration of 3-D shapes. IEEE Transactions on Pattern Analysis and Machine Intelligence, 14(2), 239–256. |
